NGS is the latest molecular-genetic examination technique that allows efficient "reading" of a large number of genetic sequences (DNA segments) simultaneously. It is a method with great potential for further development and wide application in all types of genetic examinations (targeted sequences/genes and the whole genome), including the possibility of combining them in one experiment. The NGS method (with so-called low coverage) can also be used for preimplantation genetic screening of chromosomal numerical changes (PGS (PGT-A)) or for preimplantation genetic testing of unbalanced forms of familial chromosomal rearrangements (PGT-SR) combined with screening for numerical changes of other chromosomes.
When the analysis is performed on a trophectoderm sample, it is possible, due to higher sensitivity of quantification, improved dynamic range of changes, and better signal-to-background resolution, to more reliably assess the presence of mosaic changes (compared to previously performed examinations by array comparative genomic hybridization (aCGH)). The finding of a chromosomal aberration in a mosaic means that in the sample, or in the embryo from which the sample was taken, a normal (euploid) cell line and an aneuploid line are present. These changes are always associated with de novo chromosomal aberrations detected during PGS (PGT-A) testing. In a situation where there is no (remaining) embryo with a normal finding, the transfer of an embryo with a mosaic aberration may be considered after appropriate genetic consultation and signing of informed consent.
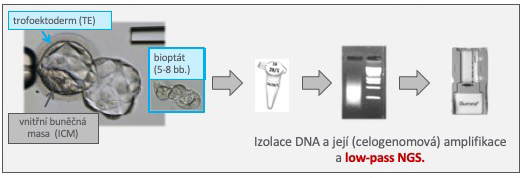
Principle of the NGS Method: DNA is extracted from the collected cells, and whole genome amplification (WGA) is then performed. Successfully amplified samples are enzymatically prepared into so-called "libraries," suitable for simultaneous "reading" of many DNA sequences. Each sample/library has its unique labeling, which allows the DNA of multiple individuals (embryos) to be analyzed in one experiment. The samples/libraries are mixed in equal proportions, and the resulting library is then prepared for final sequencing. The sequencing itself involves the step-by-step synthesis of new complementary DNA strands to the read fragments. The read sequences are compared with the normal human genome using special software, and their genomic position is determined. After assigning sequences to individual samples, a quantitative evaluation of all chromosomes is performed.
Limitations of the NGS Method: The NGS method used with low coverage (low-pass whole genome sequencing) is limited by the size of chromosomal rearrangements. It cannot detect small losses or gains of chromosomes and cannot exclude any other diseases or developmental defects of the fetus that are not caused by changes in the number of examined chromosomes or their larger parts.
Schema of NGS Examination – PGT-A, PGT-SR